Multispecific antibodies are too structurally diverse for static, stem-based naming systems. A building-block naming standard captures both the parts and the assembly logic, improving collaboration today while generating the structured data needed for AI-ready protein engineering.
Key Takeaways
Importance of a flexible, building-block naming system for MsAbs.
Need for precision and flexibility in naming to support collaborative R&D and AI readiness.
Benefits of BioGlyph Luma in visually designing and registering complex biologics.
How a building-block-based naming standard helps researchers capture entity complexity, optimize collaborative innovation, and become AI-ready for protein engineering.
Why Nomenclature Matters
Nomenclature standards may not immediately seem like the most buzzworthy topic, but they can have huge ramifications on many of today’s most important topics, including therapeutic modality innovation, research efficiency, and AI implementation. An ideal naming system should provide both ease-of-use and an extensible universal communication language for researchers as they collaboratively design, test, and communicate about candidate therapeutic entities; it must also support the highly precise data capture that is needed to not only define and differentiate complex structures, but to also enable machine-readiness for ML/AI-based work. This blog highlights key challenges of applying naming standards for dynamic next-generation biologics, like multispecific antibodies (MsAbs), and explores why solutions that leverage a building-block-based naming standard are the ideal way to support today’s workflows, while also preparing for future innovation.
Defining Multispecifics:
A Balancing Act of Precision and Flexibility
While the small molecule chemical compounds that long dominated the market lend themselves to simple, structured, stem-based naming conventions, non-natural biologics are a whole different story. Multispecific antibodies, which can bind to more than one biological target or different binding sites on the same target, are particularly challenging due to their complexity and variability. As such, any naming system for dynamic next-generation biologics, like MsAbs, must strike a balance between precision and flexibility.
Precision
Multispecific antibodies (MsAbs) are not only bigger than small molecule chemical compounds, but they are also more complex, with multiple binding regions and several moving pieces that can come together in different ways. As touched upon in the blog Managing the Madness: Three Must-Dos for Tracking Multiformat Antibody R&D, precision is essential to naming and tracking MsAbs throughout the R&D process. Not only do researchers need to systematically define the various Building Blocks (e.g., Fab, scFV, VHH, Cytokine/Mutein, Mini Protein, De Novo Protein), but they also need to be able to capture the nuances of how those Building Blocks come together in different ways to form unique entities that may have very different performance profiles. Capturing an exacting level of detail not only helps teams more easily and collaboratively iterate on complex entity design within their existing projects, but it also helps them amass the data needed to train models for in silico work, such as predictive developability assessment or De Novo design.
Flexibility
As seen in recent years, the diversity of therapeutic biologics is growing. As teams begin to pursue next-generation modalities, like MsAbs,, antibody-drug conjugates (ADCs), and gene and cell therapies, it has become obvious that static biologic design and registration solutions, which are built around predefined options, do not suffice. Many existing biologics discovery solutions struggle to accommodate the increasing complexity and diversity of biologic formats. These rigid platforms rely on imprecise representations of molecular formats, limiting flexibility and stifling the exploration of innovative biologic solutions to address unmet medical needs. Accommodating diverse and dynamic modern biologics means supporting both standardization and flexibility at once.
A Building-Block-Based Naming System
In order to provide both the precision and flexibility needed to accommodate dynamic modern biologics like MsAbs, a system must: account for all essential protein building blocks, their components, and underlying sequences; precisely define how they come together; and relate building blocks to their corresponding biological targets. Whether these building blocks were uncovered via traditional antibody discovery or derived via AI-driven De Novo design, capturing details at this intricate level provides a foundation for clearly defining and registering entities, as well as creating visual representations that help researchers better understand complex structures and push forward their research. Moreover, defining and relating entities at the building-block level enables scientists to evolve their workflows to design novel entities in the future. Scientists have the freedom to explore new formats that live outside the norm, without the fear of breaking internal registration systems.
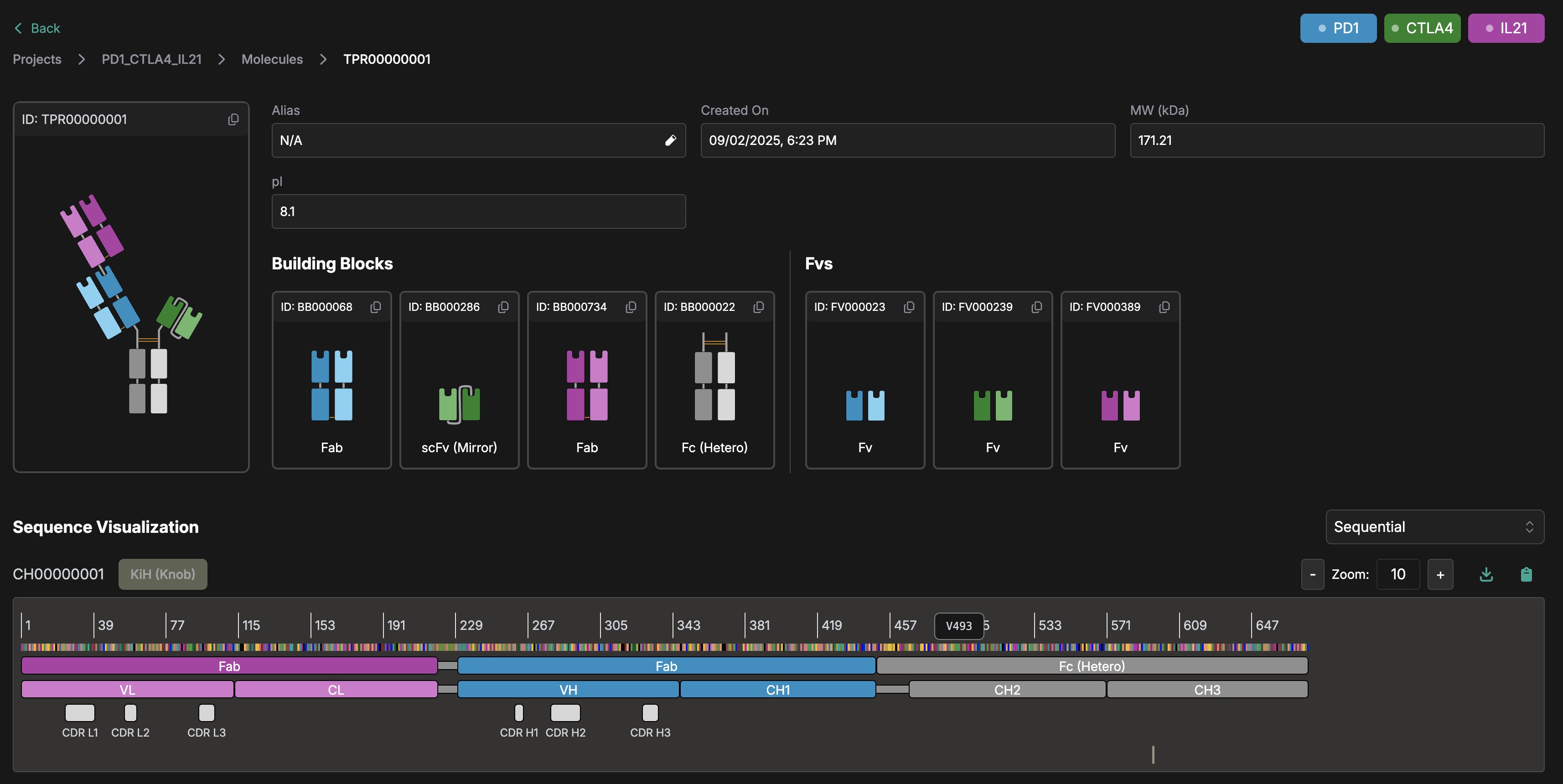
Figure: Visually defining and color-coding MsAbs at the protein-building-block level helps researchers precisely design, test, and compare various structural combinations alongside their underlying sequences.
Benefits of a Build-Block-Based Nomenclature
While there are many challenges to creating a flexible solution to design and track complex dynamic biologics, like MsAbs, one that leverages a building-block-based nomenclature provides numerous benefits, including:
Research and communication efficiency: A universal language for defining structures helps researchers more easily track and collaborate on an often overwhelming number of potential candidate structures that may have only slight design variations. And beyond internal communication, when the time comes to publish work and prepare regulatory submissions, a history of precision and consistency in defining structures across research workflows is essential.
Novel structure support: A high level of specificity not only lets researchers define and differentiate the various complex biologics structures trending in research today, but also offers the flexibility to support novel structures in the future.
AI readiness: Systematic collection of intricate structural and sequence data also serves to gather the machine-ready data needed to leverage AI/ML models, such as those for predictive developability assessment or de novo design.
Introducing BioGlyph Luma
To enable a universal language for defining complex biologics like MsAbs, a team of protein engineers and AI-expert computer engineers created BioGlyph, Inc., an agnostic and AI-powered biologics-design platform that centers around protein building blocks. BioGlyph helps researchers collaboratively design, annotate, register, and track complex biologic entities throughout the iterative design-make-test-decide R&D cycle.
Key features of BioGlyph include:
Visual interface: Design complex biologics, like MsAbs, with an intuitive drag-and-drop interface that displays molecular structures and uses color-coding to relate each component to the various targets.
Universal language: Define and register structures at the protein building-block level using BioGlyph’s proprietary, object-based language that is based on color and structure (shape).
AI predictions: Perform in silico developability assessment on the protein building blocks used to construct structures.
Dotmatics has partnered with BioGlyph to deliver AI-powered biologics-design tools from within the Dotmatics Luma platform. As part of the Dotmatics Luma Platform, BioGlyph Luma can work alongside other industry-standard software to create an end-to-end solution that optimizes both workflows and dataflows across all key steps of biologics discovery, including:
biologic entity design and engineering (BioGlyph),
cloning design and execution (Geneious),
expression, purification and PTM analysis (Protein Metrics), and
characterization and analysis (Prism statistics analysis and OMIQ/FCS Express flow cytometry analysis).
Want to see how a building-block-based approach transforms antibody format design?
Watch our on-demand webinar Unprecedented Precision in Antibody Format Design to explore BioGlyph Luma in action and learn how it supports next-gen biologics innovation.

