How can we replicate evolution in silico for proteins with unpredictable behavior?
While monoclonal antibodies (mAbs) drove the biologics revolution, a widening array of non-natural entities have entered the fold in recent years. For example, multispecific antibodies (MsAbs) are a promising way to tackle complex and hard-to-treat diseases that are not addressable with mAbs, including but not limited to cancers, inflammation,and autoimmune diseases. Researchers are looking to design non-natural biologics that attack these sorts of diseases in more than one manner. However, engineering the non-natural means thinking and working outside the norm, and as such, researchers need their biologics design and in silico prediction tools to support these novel approaches. Dotmatics and BioGlyph are working together to make AI-powered next-generation biologics design tools more accessible to researchers as they pursue novel biologic formats, from multispecifics and beyond.
Next-Generation Design Demands Next-Generation Solutions
As researchers shift their focus to next-generation biologics, they often face a key barrier to innovation: R&D tools built around rigid, predefined entity structures. Tools that may have served researchers well as they developed simpler modalities, like mAbs, are now showing their shortcomings. To engineer next-generation biologics, like MsAbs, teams need next-generation solutions. As described below, in order to promote continued biologics innovation in areas such as MsAb design, these solutions must offer a few key must-haves.
1. Computational entity design via a building-block-based approach
To design complex non-natural entities, like MsAbs, researchers need to be afforded the opportunity to work outside the norm. This isn’t possible with rigid solutions that only support pre-defined structure options and lack the capabilities to precisely define complex novel entities. On the other hand, design tools that offer a building-block-based approach, like BioGlyph Luma, can support the structural complexity of next-generation biologics; this may be MsAbs today and novel entities in the future. Researchers need to be able to define how building blocks of their entities come together (e.g., Fab, scFV, Fc, VHH, Mini Protein, de Novo proteins, Cytokines/Muteins), tie structures to their underlying sequences, and leverage visualization to more easily differentiate and scrutinize designs. Beyond this, they need to be able to easily track and iterate on different design tweaks as they work to uncover the precise designs that will maximize proper assembly and performance, and ultimately best ensure the success of antibodies produced in the lab.
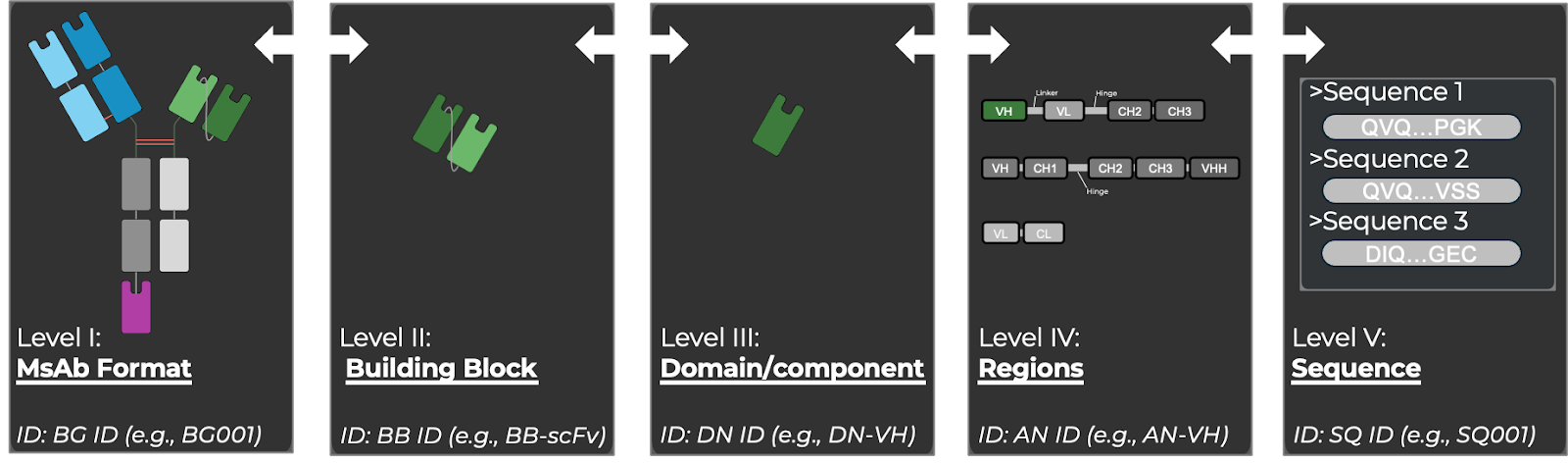
Figure: Five levels of annotation interconnecting MsAb format (quaternary structure), to Building Block, Component, Regions and single amino acids.
2. In silico assessment to select candidates brought to the lab
The structural and functional complexity of MsAbs make them prone to unpredictable behavior. In fact, MsAbs have a relatively high failure rate in clinical trials.[1] As structural complexity increases, so does the range of issues that can impact performance, such as assembly mispairings, incorrect foldings, suboptimal alignment, etc. To get ahead of these issues early on, researchers are turning to in silico tools; for example:
Rational design tools can aid in the selection of the precise building blocks that are most likely to lead to efficient assembly, functional stability, and manufacturability.
Modeling tools, such as DeepMind’s AlphaFold and the Baker Lab’s Rosetta Fold, can help researchers predict protein structures and model interactions.
De novo design tools can help experts derive novel formats that optimize structure and function, for example designing a new component to improve linker design, and thereby, MsAb stability.
However, MsAbs have more complex binding kinetics and mechanism of actions compared to their mAb counterparts. Naturally, with an increase in functional areas, there is likewise an increase in potential behaviors (and misbehaviors).Therefore, behavioral prediction is inherently complicated when dealing with structures that have been engineered to serve multiple purposes, whether that be binding to multiple targets, recruiting immune cells, or serving as molecular bridges. While most biologics design platforms on the market are too rigid to model and make predictions about complex next-generation entities, like MsAbs, BioGlyph Luma is different. Researchers can iterate on their designs to evolve initial concepts into refined candidates that have the best chances of success in the lab and beyond.
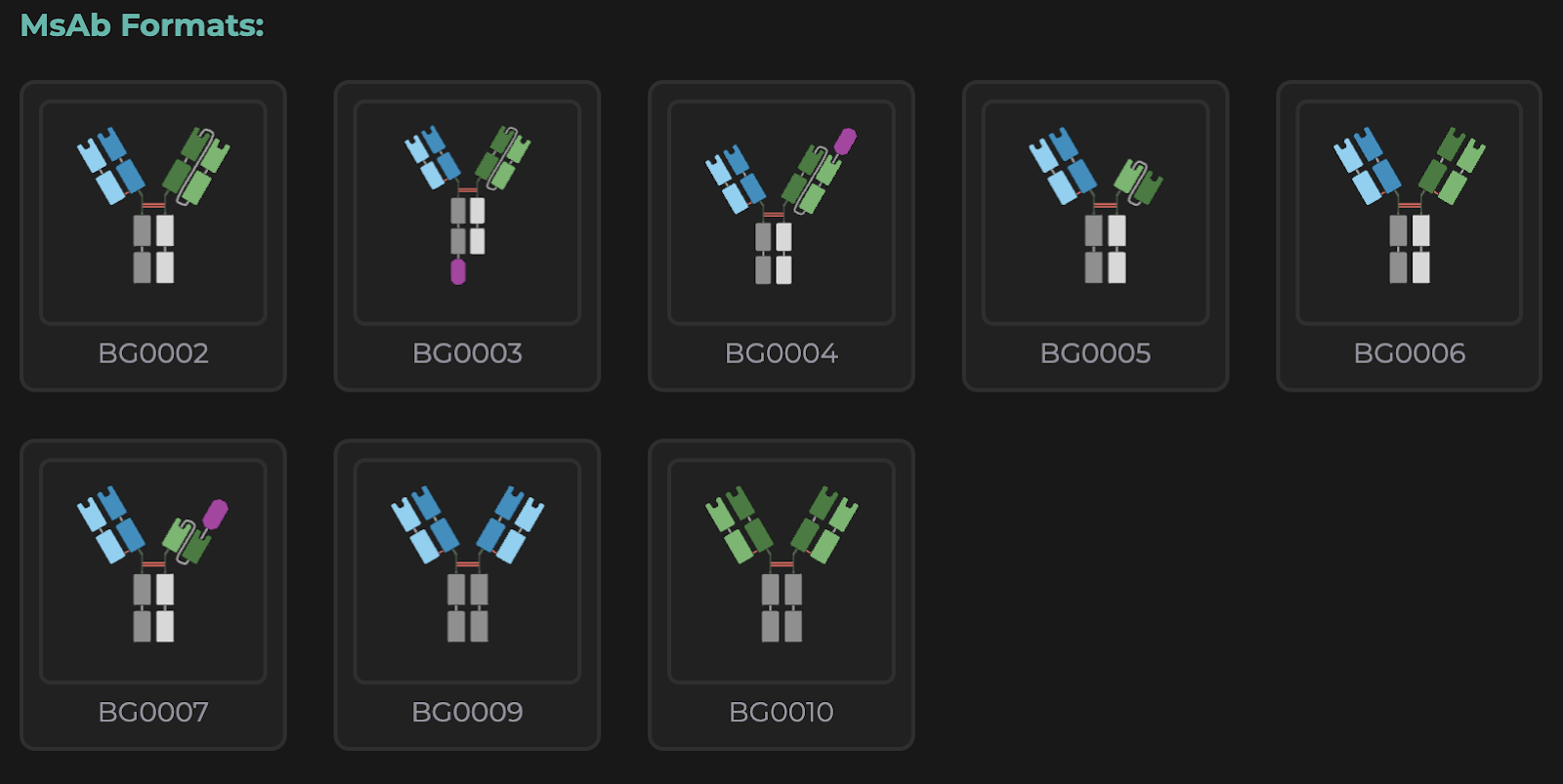
Figure: Example of MsAb formats users can design. While the naming structure is driven by color and shape, a unique ID is generated to also capture format uniqueness.
3. Lab-in-a-loop to optimize R&D cycles and propel structural evolution
As teams bring their virtually designed structures into the wet lab, they need to collect and relate data across many workflows steps—from cloning design and execution, to expression, purification and PTM analysis, to characterization and analysis. The wealth of knowledge gleaned in the web lab needs to be looped back to the virtual lab, where designs can be further evolved. This iterative process involves a wide range of experts using specialty computational design, molecular biology, sequence analysis, proteomics, and characterization and analysis tools that sometimes don’t work together well; this inevitably leads to bottlenecks, errors, and missed connections. Dotmatics is changing this paradigm by creating an end-to-end platform that enables seamless data exchange and real-time traceability across workflows in both the physical and virtual lab. With each structure assigned a unique ID, candidates can be fully traced as they are designed, produced, and tested in downstream workflows by experts using different specialty instruments and applications. This enables teams to more easily collect the data/metadata that is not only needed to collaboratively evolve entity design, but is also essential when submitting for regulatory review.
BioGlyph Luma, an integrated AI-powered next-generation biologics design tool
Dotmatics and BioGlyph have partnered to integrate AI-powered, next-generation biologics-design tools into the Dotmatics Luma scientific R&D platform. BioGlyph Luma is the latest of several specialty science applications—including Geneious, Protein Metrics, and Prism/Flow—that have been brought onto the Luma platform to help researchers optimize biologics R&D efforts across their in silico and in vitro workflows.
Reference
Daniel Keri, Matt Walker, Isha Singh, Kyle Nishikawa, Fernando Garces, Next generation of multispecific antibody engineering, Antibody Therapeutics, Volume 7, Issue 1, January 2024, Pages 37–52, https://doi.org/10.1093/abt/tbad027
